-
- Downloads
Add elements on requirements.txt
Showing
- code/bolsonaro/visualization/plotter.py 12 additions, 14 deletionscode/bolsonaro/visualization/plotter.py
- code/compute_results.py 15 additions, 0 deletionscode/compute_results.py
- requirements.txt 2 additions, 1 deletionrequirements.txt
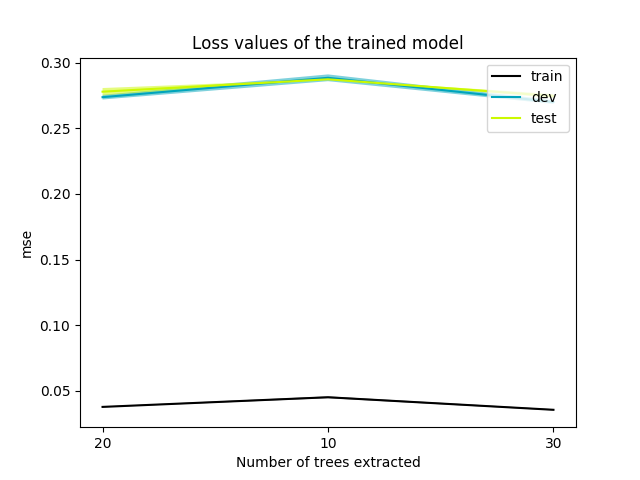
- results/1/losses.png 0 additions, 0 deletionsresults/1/losses.png
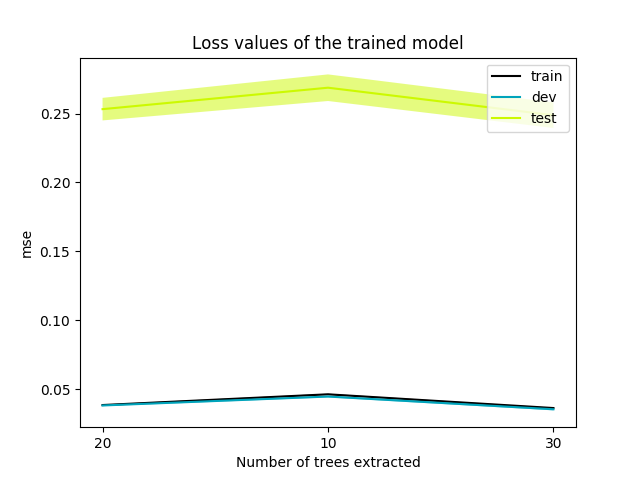
- results/2/losses.png 0 additions, 0 deletionsresults/2/losses.png
| ... | ... | @@ -12,3 +12,4 @@ scikit-learn |
| python-dotenv | ||
| tqdm | ||
| matplotlib | ||
| pandas | ||
| \ No newline at end of file |
results/1/losses.png
0 → 100644
24.7 KiB
results/2/losses.png
0 → 100644
24 KiB